Advanced Deep Convolutional Neural Network
Advanced Deep Convolutional Neural Network Approaches for Digital Pathology Image Analysis
Introduction
Medical imaging speeds up the assessment process of almost every disease, from lung cancer to heart disease. The automatic pathological image classification, segmentation, and detection algorithm can help to unlock the cure faster from critical diseases, like cancer to common cold. The computational pathology and microscopy images play a big role in decision making for disease diagnosis. Therefore, this solution can help to ensure better treatment. Nowadays, there are different DCNN models that have been successfully applied in computational pathology. We have applied three different improved DCNN models for pathological image classification, segmentation, and detection. The overall implementation diagram is shown in Figure 1.
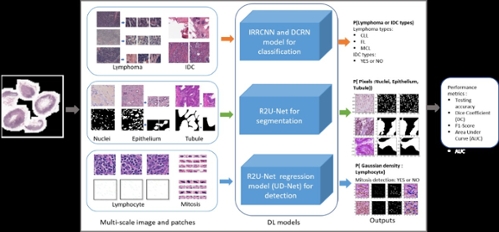 Figure 1. Overall deep learning framework for computational pathology in seven different tasks.
Figure 1. Overall deep learning framework for computational pathology in seven different tasks.
Goals/Objectives
We have proposed new DCNN models (IRRCNN: Inception Residual Recurrent CNN, DCRN: Densely Connected Recurrent Network) for Lymphoma, Invasive Ductal Carcinoma (IDC), and mitosis classification.
To generalize the Recurrent Residual U-Net (R2U-Net) model, R2U-Net is applied for nuclei segmentation, epithelium segmentation, tubule segmentation.
The UD-Net is proposed for end-to-end lymphocyte detection from pathological images.
The experimental results show superior performance compared to existing machine learning and DL based approaches for classification, segmentation, and detection tasks.
Methods
We have applied these advanced DCNN techniques including IRRCNN, R2U-Net, DCRN (an improved version of DenseNet), and a R2U-Net based regression model called University of Dayton Net (UD-Net). UD-Net uses a cell image I(x) as input and computes the density heat maps D(x). Figure 2 shows training and validation accuracy for IRRCNN and DCRN.
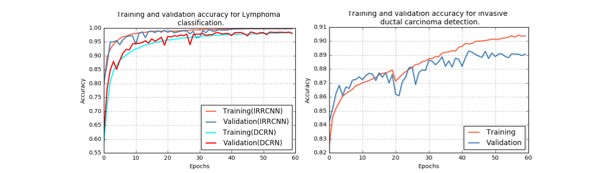 Figure 2. The training and validation accuracy for Lymphoma classification with IRRCNN and DRCN on the left and for invasive ductal carcinoma classification on the right.
Figure 2. The training and validation accuracy for Lymphoma classification with IRRCNN and DRCN on the left and for invasive ductal carcinoma classification on the right.
Table 1: The quantitative results and comparison against existing approaches for seven different tasks.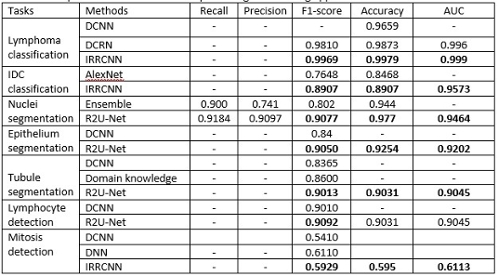
Results/Evaluations
We have evaluated the performance of these models with different metrics such as precision, recall, accuracy, F1-score, area under the receiver operating characteristics curve (AUC), dice coefficient (DC), and mean squared error (MSE). The quantitative results are shown in Table 1. It can be seen that the proposed methods provide better performance on different tasks. The qualitative results are shown in Figure 3.
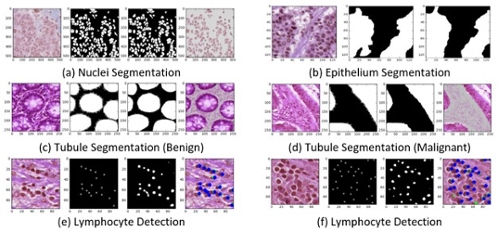 Figure 3. Experimental results with R2U-Net. The first column shows the input images, the second column shows the ground truth (GT), the third column shows the model outputs, and the fourth column shows the overlapped target regions.
Figure 3. Experimental results with R2U-Net. The first column shows the input images, the second column shows the ground truth (GT), the third column shows the model outputs, and the fourth column shows the overlapped target regions.
Paper link: https://arxiv.org/ftp/arxiv/papers/1904/1904.09075.pdf
